Mechanism of action of enzyme for complex natural product biosynthesis Toward enzymatic regulation of indole prenylation

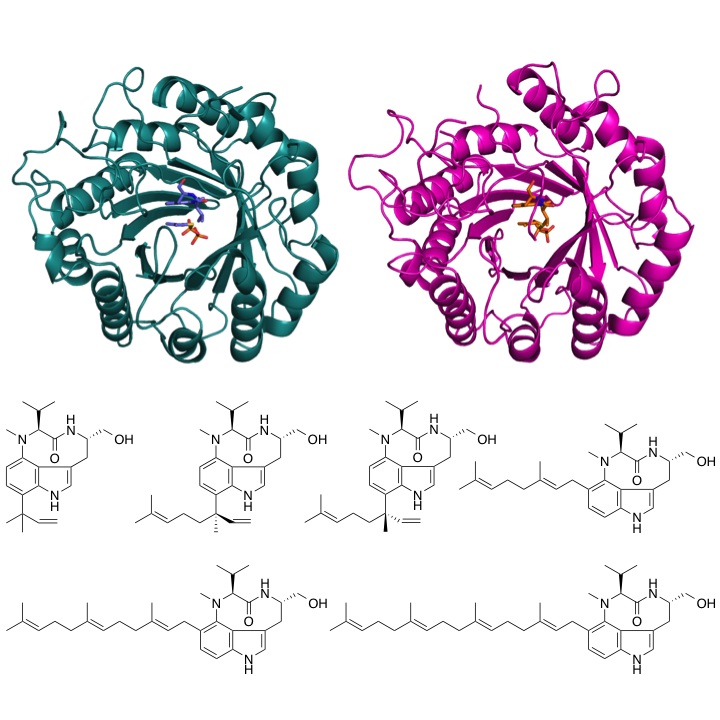
Crystal structure of enzymes biosynthesizing terpene indole alkaloids
The two prenyltransferase transfer prenyl substrates with a range of chain lengths to a specific position on the indole ring.
© 2016 Ikuro Abe.
Researchers at the University of Tokyo have revealed the structure and enzymatic mechanism of a prenyltransferase, an enzyme involved in the biosynthesis of complex natural products. The results pave the way for enzymatic design of novel molecular scaffolds, contributing to medicinal chemistry.
Natural products, the chemical compounds produced by living organisms, have immense structural diversity and are a valuable source for drug discovery. Among them are the terpene indole alkaloids, a group of compounds that fulfill a wide variety of biological functions. They are biosynthesized by addition of prenyl groups, chains formed of multiples of five carbon atoms, to indoles, compounds that we sense as smelling rancid, creating intermediates called prenylated indoles. The enzymes responsible for the biosynthesis of the terpene indole alkaloids within living organisms are thus of interest for the development of biocatalysts with unique activities. However, much remained unknown about the reaction mechanism by which the enzyme added the prenyl group to the indole compound.
The research group of Assistant Professor Takahiro Mori and Professor Ikuro Abe at the University of Tokyo Graduate School of Pharmaceutical Sciences has obtained two crystal structures of the enzyme that catalyzes prenylation of prenyl donors to indole substrates of varying length. Based on the crystal structure analysis, they successfully revealed that the enzyme employs a “reverse” prenylation mechanism, the opposite of canonical prenylation reactions. Furthermore, by comparing the two crystal structures, the research group identified the molecular mechanism of substrate recognition by which the enzyme distinguishes the length of prenyl chain, and showed that the enzymes can accept prenyl substrates of up to 25 carbon atoms in length, which is the longest substrate accepted by the same class of enzymes.
“The ability to catalyze a prenylation reaction of such a long carbon chain to an indole is an unusual characteristic for an indole prenyltransferase,” says Assistant Professor Mori. He continues, “Prenylation is known to be an important step to improve bioactivity of compounds, and our results will pave the way for the development of biocatalysts that can generate novel molecular scaffold for drug discovery, catalyzing reactions difficult to achieve by organic synthesis.”
Paper
, "Manipulation of prenylation reactions by structure-based engineering of bacterial indolactam prenyltransferases", Nature Communications Online Edition: 2016/03/08 (Japan time), doi: 10.1038/ncomms10849.
Article link (Publication)
Links
Graduate School of Pharmaceutical Sciences
Laboratory of Natural Product Chemistry, Graduate School of Pharmaceutical Sciences






