DNA methylation switches in evolution Evidence from whole methylome decoding in Helicobacter pylori
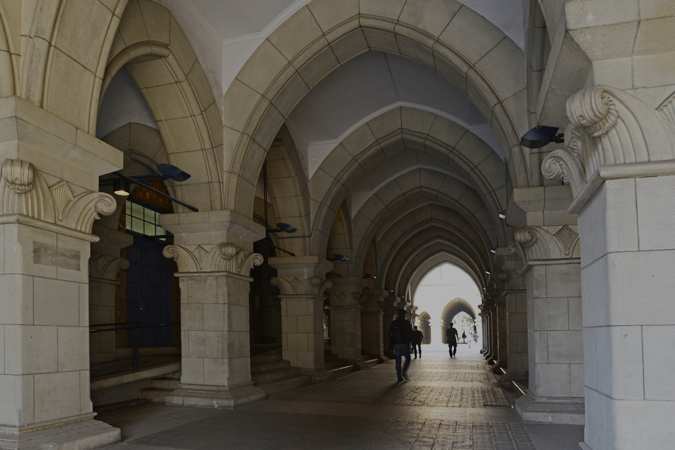

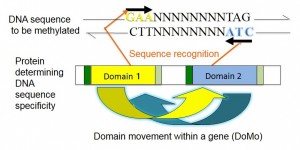
Recognition of DNA sequence to be methylated by a protein. The protein comes to recognize a different sequence when intragenic recombination moves an amino acid sequence between two domain sites.
CC0 1.0
The authors dedicate this work to the public domain by waiving all of his or her rights to the work worldwide under copyright law, including all related and neighboring rights, to the extent allowed by law.
It is accepted in the mainstream of life sciences today that the basic unit of heredity is to be found in the genetic information written in the sequence of nucleotides in DNA. However, even when the nucleotide sequence remains unchanged, if a specific nucleotide sequence is modified by means such as methylation, the state of the DNA (epigenetic state) changes and can lead to genes being expressed or suppressed. This epigenetic information can be passed on through genomic replication, a fact that is currently attracting attention. As a result it is possible that the real unit of heredity is to be found in this epigenetic information overwritten on the DNA nucleotide sequence.
Professor Ichizo Kobayashi and Project Assistant Professor Yoshikazu Furuta of the University of Tokyo Graduate School of Frontier Sciences and their colleagues revealed the methylation state of each individual nucleotide in the entire genomes of multiple strains of the bacteria Helicobacter pylori. The researchers found that rearrangements within a gene that specifies sequence specificity of methylation changed the methylation pattern. When such a gene was deleted, gene expression pattern was altered.
The present study is consistent with the hypothesis that epigenetic information is important in the evolution of living organisms, and is expected to pave the way for “epigenomic breeding” to artificially manipulate the epigenome.
Paper
Yoshikazu Furuta, Hiroe Namba-Fukuyo, Tomoko F. Shibata, Tomoaki Nishiyama, Shuji Shigenobu, Yutaka Suzuki, Sumio Sugano, Mitsuyasu Hasebe, Ichizo Kobayashi,
“Methylome Diversification through Changes in DNA Methyltransferase Sequence Specificity”,
PLoS Genetics Online Edition: 2014/4/10, doi: 10.1371/journal.pgen.1004272.
Article link
Links
Graduate School of Frontier Sciences
Department of Medical Genome Sciences, Graduate School of Frontier Sciences
Kobayashi Laboratory, Department of Medical Genome Sciences, Graduate School of Frontier Sciences