Hybrid genes related to pediatric leukemia with poor prognosis discovered Researchers identify fusion genes in pediatric T-cell lymphoblastic leukemia
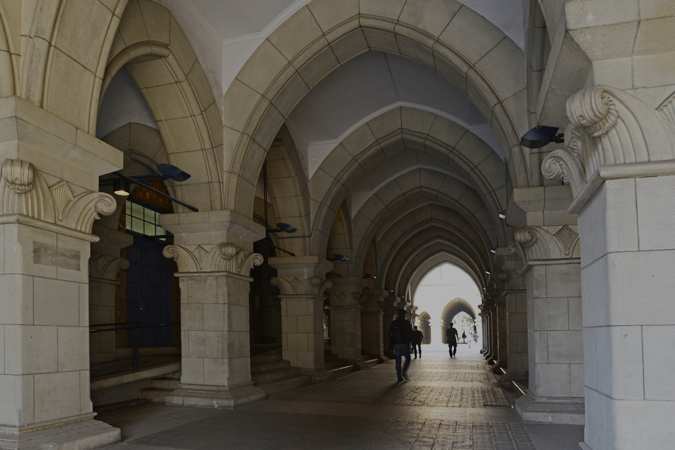

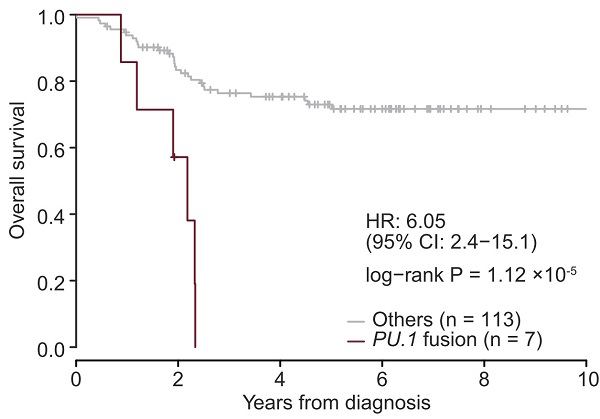
Overall survival in T-ALL cases with, without SPI1 fusion genes
When comparing cases of pediatric T-cell acute lymphoblastic leukemia (T-ALL), a type of blood cancer afflicting children, those involving SPI1 fusion genes (represented as PU.1 fusion above) show significantly shorter overall survival than cases lacking the hybrid gene, with six in seven patients dying within 3 years of diagnosis.
© 2017 Masafumi Seki.
Researchers at the University of Tokyo and their collaborators have identified recurrent fusion genes—new genes formed from the combination of two or more separate genes that appear in more than one sample of a disease taken from different patients—tied to a form of pediatric T-cell acute lymphoblastic leukemia (T-ALL), a type of blood cancer occurring in a subtype of white blood cells, with an extremely poor prognosis; moreover, the team demonstrated that the genes' genetic and clinical characteristics differ greatly from other T-ALL cases. The study's findings hold promise of leading to more precise therapies and optimal treatments for the disease.
Recent advances in treatment, incorporating intensive chemotherapy combined with other treatments, have led to about a 70 percent cure rate for pediatric T-ALL, which comprises roughly 15 percent of all childhood leukemia. However, late effects, the health conditions occurring after treatment, such as growth failure, organ dysfunction, and infertility, remain to be serious problems affecting survivors. Moreover, the outcome for patients with T-ALL that is primary resistant (not responding to treatment from the outset) or which has relapsed remains extremely poor. For these reasons, an optimal treatment strategy based on molecular biology is imperative for improving survival and preventing severe side effects and late effects in pediatric T-ALL.
The research group led by Associate Professor Junko Takita, Assistant Professor Masafumi Seki, and Dr. Shunsuke Kimura at the Department of Pediatrics at the University of Tokyo Hospital produced comprehensive profiles of genetic abnormalities, fusion genes, and structural variants in the genomes (the complete genetic information) of 123 cases of pediatric T-ALL using next-generation sequencing, which can determine the order of DNA bases much faster than previous gene sequencers. The researchers identified new recurrent fusion genes, containing a segment of a gene called SPI1, related to extremely poor prognosis in 4 percent of pediatric T-ALL. SPI1 is a transcription factor, a type of protein responsible for regulating the transcription of genetic information, instrumental in the differentiation of blood cells; diminished expression of SPI1 in immune cells known as T cells, a subtype of white blood cells, is thought to play an important role in differentiation. The group demonstrated that overexpression of SPI1 fusion genes blocks the differentiation and proliferation of T cells, thereby pointing to the possibility that this induces the development of leukemia. Furthermore, it revealed by profiling gene expression and examining molecular features of pediatric T-ALL that the disease could be divided into five clusters, each characterized by specific gene expression, genetic abnormalities, and clinical features.
The current study shows that the newly discovered recurrent SPI1 fusions, exhibiting unique cytological and gene-expression profiles, are a distinct subgroup of pediatric T-ALL. The contribution of these findings to T-ALL prognosis, the development of highly accurate molecular diagnosis, and optimized treatment looks promising.
"There hadn't been any reports so far on molecular markers related to pediatric T-ALL with a poor prognosis," says Takita. She continues, "Cases with SPI1 fusion genes show a dismal prognosis, likely requiring a higher risk classification and more intensive treatment. Targeted therapy employing highly accurate molecular diagnosis could prove effective in treating the disease in the future."
This research outcome is the result of a collaboration with Professor Seishi Ogawa at the Department of Pathology and Tumor Biology, Graduate School of Medicine, Kyoto University, and other researchers.
Press release (Japanese)
Paper
, "Recurrent SPI1 (PU.1) fusions in high-risk pediatric T cell acute lymphoblastic leukemia", Nature Genetics Online Edition: 2017/07/04 (Japan time), doi: 10.1038/ng.3900.
Article link (Publication)
Links
Department of Pediatrics, University of Tokyo Hospital (Japanese)