Chemical modification in RNA changes genetic code Novel transfer RNA modification establishes non-universal genetic code

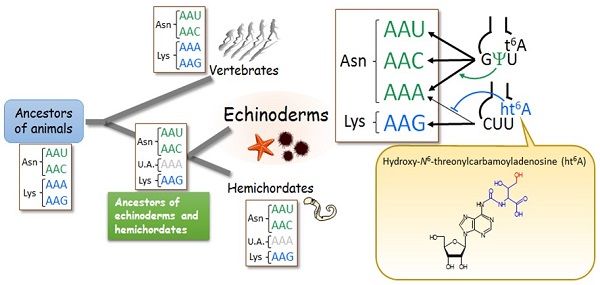
AAA codons reassigned by transfer RNA modifications in echinoderms
Scientists believe the AAA codon is unassigned (U.A.)—i.e., a genetic code that is not translated into a specific amino acid—in the common ancestors of echinoderms, including sea urchins and starfish, and hemichordates, which are wormlike marine invertebrates. In echinoderms, the AAA codon was captured by tRNAAsn (with a GΨU anticodon). A modification, ht6A37, prevents tRNALys from misreading AAA as lysine; thus, the AAA codon has switched completely, coding for lysine instead of asparagine.
© 2018 Asuteka Nagao and Tsutomu Suzuki.
A University of Tokyo research group discovered a novel modified base in one of the transfer RNA of echinoderm mitochondria, and has shown this modification plays a critical role in decoding a codon that deviates from the universal genetic code.
The so-called universal genetic code discovered in the 1960s was considered common to all living organisms and responsible for converting the genetic information contained in DNA to proteins, with all living things sharing the same code. This apparent conservation of the genetic code assumes that all domains of life evolved from a single interbreeding organism. However, genetic codes deviating from this universal code are frequently found in the genetic translation system in animal mitochondria, the organelles that serve as the powerhouse for generating energy in animal cells, raising an alternative theory that the genetic code is not fixed, but instead continues to evolve. Post-transcriptional modification of transfer RNA (tRNA) is thought to play a critical role in establishing such non-universal genetic codes. In the mitochondrial decoding system of echinoderms, including organisms like sea urchins and starfish, the AAA codon specifies the amino acid asparagine (Asn) instead of lysine (Lys). A previous study revealed that tRNAAsn, the tRNA carrying asparagine, acquires pseudouridine (Ψ) at the second letter of the anticodon, and reads the AAA codon as asparagine. However, the mechanism by which tRNALys, the tRNA carrying lysine, prevents misreading the AAA codon as lysine remained elusive.
The research group led by Assistant Professor Asuteka Nagao and Professor Tsutomu Suzuki of the Department of Chemistry and Biotechnology at the University of Tokyo's Graduate School of Engineering analyzed mitochondrial tRNALys isolated from the Mesocentrotus nudus sea urchin, and discovered a novel modified base, hydroxyl-N6-threonylcarbamoyladenosine (ht6A), in a part adjacent to the anticodon. The researchers examined the codon recognition ability of this modification on the ribosome, the apparatus for protein synthesis in the cell, and found that it is capable of preventing tRNALys from misreading the AAA codon as lysine. These results suggest that tRNALys acquired a novel modification that contributes to the genetic code alteration of the AAA codon in echinoderm mitochondria.
This discovery not only adds new knowledge for RNA research, but also suggests that the establishment of a non-universal genetic code relies heavily on the chemical modification of tRNA.
"'There is a modified base with a molecular mass that we have never seen!' Such was our response to our exciting adventure of discovering the new modification with just the 16-dalton mass difference as our clue, and studying its molecular function," says Nagao. He continues, "That one oxygen atom governs the genetic code alteration was very interesting from both biochemical and evolutionary points of view. I would like to continue investigating a mechanism for building life that has not been discovered yet."
Paper
, "Hydroxylation of a conserved tRNA modification establishes non-universal genetic code in echinoderm mitochondria", Nature Structural & Molecular Biology Online Edition: 2017/08/07 (Japan time), doi: 10.1038/nsmb.3449.
Article link (Publication)
Links
Graduate School of Engineering
Department of Chemistry and Biotechnology, Graduate School of Engineering
Suzuki Laboratory, Department of Chemistry and Biotechnology, Graduate School of Engineering






