First step to deciphering a non-universal genetic code Identification of an enzyme required for reading mitochondrial genome

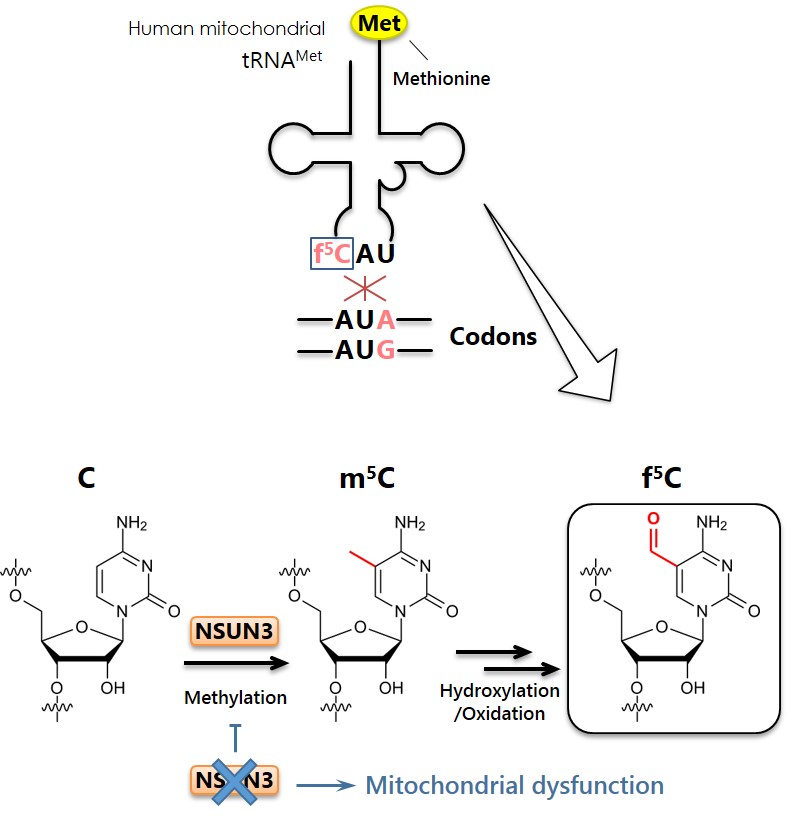
Human mitochondrial methionine tRNA and f5C modification
(Right) The f5C modification in the anticodon enables mitochondrial tRNA (tRNAMet) to read the AUA codon in addition to the AUG codon as methionine. (Left) A putative biogenesis of f5C modification. First, the newly-discovered enzyme NSUN3 methylates cytidine (C) to form 5-methylcytidine (m5C). Then, m5C is further hydroxylated and then oxidized to form f5C. A lack of the NSUN3 enzyme results in lack of the f5C modification and the inability to read the AUA codon results in reduced mitochondrial function.
© 2016 Tsutomu Suzuki
A University of Tokyo research group has discovered a novel enzyme that introduces a transfer RNA (tRNA) modification required to decipher a non-standard genetic code in the DNA of human mitochondria. The group showed that lack of this enzyme or of the tRNA modification results in mitochondrial dysfunction and disorders.
Mitochondria are small organelles found inside cells that generate most of the cellular energy and are essential to sustaining cell life and health. Mitochondria have their own genome that contains the genes for creating mitochondrial proteins. Like genes in the cell’s nucleus, genes in the mitochondrial genome are composed of groups of three nucleotides called codons, specifying the sequence of amino acids in the protein. In translation, the process of creating proteins from genes, tRNA molecules that recognize a particular codon carry a single amino acid to the protein manufacturing machinery, where the amino acids are attached one after the other to create proteins. The codons are recognized by a section of the tRNA molecule called an anticodon. Although a universal genetic code dictates which codon matches which amino acid, the adenine-uracil-adenine (AUA) codon that normally specifies the amino acid isoleucine specifies methionine in human mitochondria. It is known that this different code requires a tRNA with a modified anticodon containing 5-formylcytidine (f5C). However, how this anticodon modification is produced and its physiological role remain unknown.
The research group of then-graduate student Saori Nakano, Lecturer Takeo Suzuki, Professor Tsutomu Suzuki and their colleagues at the Department of Chemistry and Biotechnology, Graduate School of Engineering found that human cells lacking the NSUN3 gene also lacked the f5C modification in the mitochondrial tRNA for methionine, and observed severe dysfunctions of mitochondrial activity in these cells. In addition, they showed that the protein NSUN3 encoded by this gene methylated the anticodon of the tRNA to form an intermediate, which they suggest is then oxidized by an unknown enzyme to form f5C.
Several mutations identified as causing mitochondrial diseases have been reported in the gene for mitochondrial tRNA that carries methionine. The research group found that two of these mutations greatly impaired the formation of the m5C modification by NSUN3.
“It was a tough challenge to identify this enzyme. But the fact that this modification is essential for human health and lack is associated with some diseases was a strong motivation to continue until we found it,” says Nakano. She continues, “I clearly remember the moment when I found a lack of f5C in the knockout cells. I felt my efforts were rewarded.”
“Little was known about the biogenesis and physiological role of f5C, even though it was discovered more than 20 years ago,” says Takeo Suzuki. He has great expectations of further progress of their research commenting, “We still need to unveil the second step of f5C biogenesis after methylation. Considering that the loss of f5C is associated with human disease, I hope our study will contribute to medical applications.”
Paper
, "NSUN3 methylase initiates 5-formylcytidine biogenesis in human mitochondrial tRNAMet", Nature Chemical Biology: 2016/5/24 (Japan time), doi: 10.1038/nchembio.2099.
Article link (Publication)
Links
Graduate School of Engineering
Department of Chemistry and Biotechnology, Graduate School of Engineering
Suzuki Laboratory, Department of Chemistry and Biotechnology, Graduate School of Engineering






