Mechanism by which a protein binds a target Intrinsically disordered proteins bind partners by two mechanisms

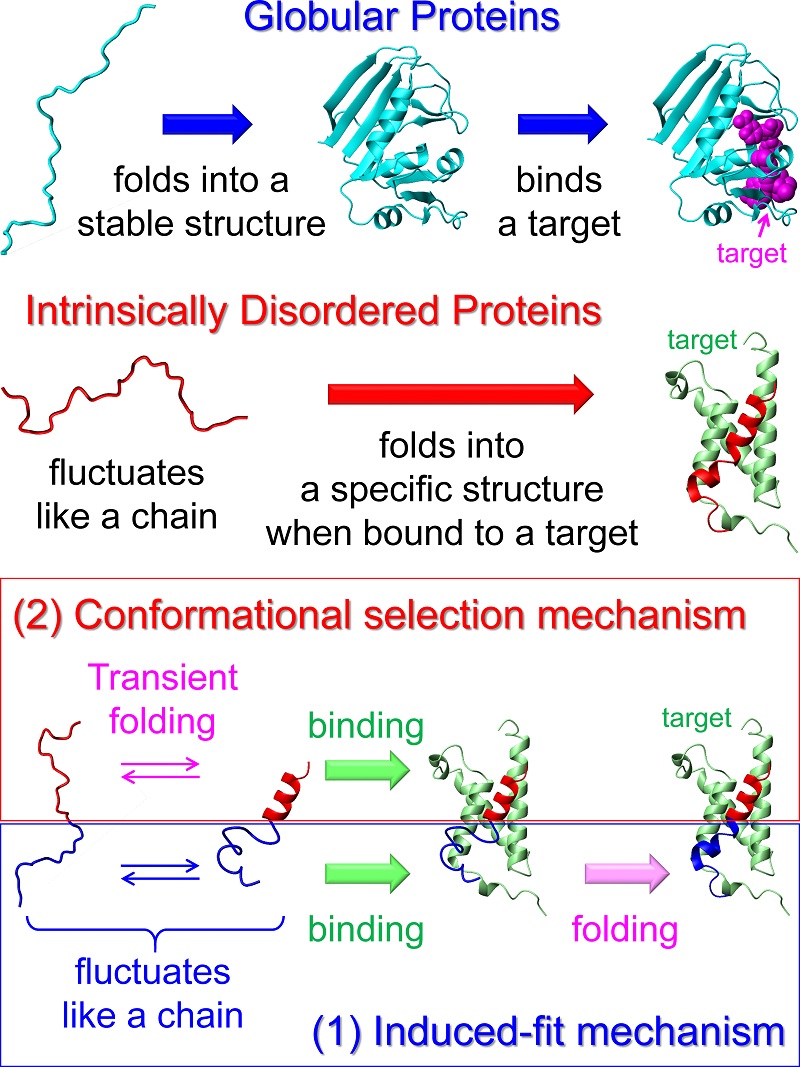
Mechanism by which an IDP binds a target molecule
Two major mechanisms have been proposed for IDP folding upon binding. One is an induced-fit mechanism, in which binding precedes folding. Another is a conformational selection mechanism, in which an IDP binds its target only when it transiently folds into a specific structure. In the case of a typical IDP c-Myb, its upper region (red) binds a target protein KIX (green) by the conformational selection mechanism, while its lower region (blue) binds KIX by the induced-fit mechanism. c-Myb is a transcription factor that facilitates proliferation of hematopoietic cells and is associated with leukemia.
© 2015 Munehito Arai.
Researchers at the University of Tokyo and the Scripps Research Institute have elucidated the mechanism by which an intrinsically disordered protein (IDP), a novel class of proteins, binds a target molecule. The results may help elucidate how disease-related IDPs work and help drug design.
Many proteins perform their functions in living organisms through binding to specific partners. A protein is a chain-like molecule and, in many cases, it binds to its partner after folding into a stable structure. On the other hand, an IDP is disordered in isolation, but folds into a specific structure when bound to its target. However, the mechanism of IDP folding upon binding was poorly understood.
In collaboration with the research group of Professor Peter E. Wright (The Scripps Research Institute, USA), Associate Professor Munehito Arai (Graduate School of Arts and Sciences, The University of Tokyo) has measured the binding reaction of a typical IDP with its target molecule using nuclear magnetic resonance spectroscopy. They found that IDPs bind their target molecules by two mechanisms, induced-fit and conformational selection, depending on their conformational propensities.
Many proteins related to diseases such as cancer, leukemia, Alzheimer’s disease, Parkinson’s disease, and bovine spongiform encephalopathy are categorized as IDPs. The present results may help elucidate how these proteins perform their functions and may help develop drugs for the diseases.
Press release (Japanese)
Paper
, "Conformational propensities of intrinsically disordered proteins influence the mechanism of binding and folding", Proceedings of the National Academy of Sciences of the United States of America vol. 112, no. 31, page 9614-9619, doi: 10.1073/pnas.1512799112.
Article link (Publication)
Links
Graduate School of Arts and Sciences
Department of Life Sciences, Graduate School of Arts and Sciences (Japanese)
Arai Lab, Department of Life Sciences, Graduate School of Arts and Sciences (Japanese)






