CO2-sensitive RNA modification Possible link between tRNA modification and mitochondrial regulation in cancer cells
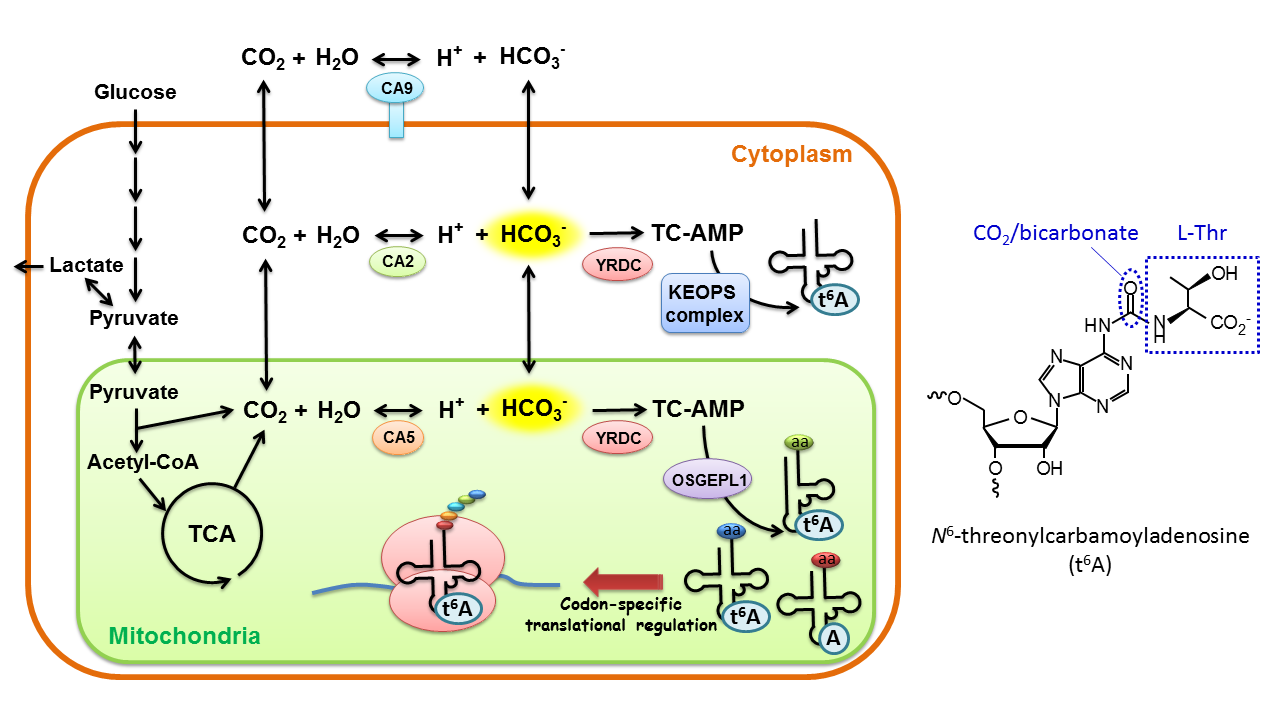
Carbon dioxide (CO2) is produced by mitochondria during respiration. CO2 penetrates cellular membranes as a gas. CO2 is hydrated to generate bicarbonate (HCO3-) by the action of carbonic anhydrase (CA). Under hypoxic conditions, CA9, an isoform of CA, is overexpressed on the plasma membrane and produces a large amount of bicarbonate to neutralize lactate produced by anaerobic glycolysis, thereby preventing acidosis. This study revealed that bicarbonate is used as a substrate for t6A biogenesis and plays a critical role in tRNA function. Upon depletion of CO2 or HCO3- in the cell, t6A frequency in mitochondrial tRNAs is reduced and mitochondrial protein synthesis is regulated in a codon-specific manner.
© 2018 Tsutomu Suzuki
A University of Tokyo research group discovered that transfer RNA modification is dynamically regulated by sensing either cellular carbon dioxide or bicarbonate concentration in human mitochondria.
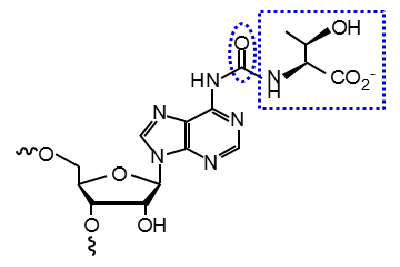
N6-threonylcarbamoyldenosine (t6A) is a transfer RNA (tRNA) modification in which the amino acid L-threonine (L-Thr) is conjugated with the N6-amino group of the adenine base through a carbon dioxide (CO2)-derived carbonyl group. t6A is present at position 37, which is 3’adjacent to the anticodon, and plays critical roles in various stages of protein synthesis. t6A is a highly conserved modification found in all organisms from bacteria to humans, which is required for cells to grow efficiently.
The research group led by graduate student Huan Lin and Professor Tsutomu Suzuki of the Department of Chemistry and Biotechnology at the Graduate School of Engineering, the University of Tokyo, identified two enzymes, YRDC and OSGEPL1, responsible for biogenesis of t6A in human mitochondrial tRNA.
In fact, the researchers found that t6A disappeared when the OSGEPL1 gene was knocked out, and that OSGEPL1 knockout cells showed marked reduction of protein synthesis and mitochondrial dysfunction. Then, they successfully reconstituted t6A formation in vitro using these recombinant proteins in the presence of four substrates including L-Thr, bicarbonate (HCO3-), ATP and tRNA.
Kinetic measurement of t6A formation revealed that the Km value of HCO3- was extremely high (31 mM). Considering that bicarbonate concentration in mitochondria ranges from 10 to 40 mM, HCO3- is a rate-limiting factor for t6A formation, raising the possibility that t6A is dynamically regulated under physiological conditions. Consistent with this, the researchers observed a low frequency of t6A in mitochondrial tRNAs isolated from human cells cultured under a low bicarbonate concentration. Although tRNA modifications are generally regarded as stable and static, this result demonstrated that t6A is dynamically regulated by sensing bicarbonate concentration. Since low t6A frequency directly reduces the decoding ability of tRNA, the regulation of mitochondrial translation under some environmental or respiratory conditions is likely.
This study provides a novel concept of regulatory gene expression mediated by dynamic alteration of tRNA modification by sensing cellular metabolic status. Most cancer cells reduce mitochondrial activity and increase anaerobic glycolysis. This is called the Warburg effect. Given that most cellular CO2 is produced during respiration, CO2 sensitivity to t6A might be a key mechanism for shifting from respiration to anaerobic glycolysis by slowing down mitochondrial translation.
“I was very excited to find out that the most widely known and essential tRNA modification is regulated by a simple metabolite, CO2 gas, challenging the conventional wisdom that tRNA modifications are stable and static,” says Dr. Lin. He continues, “This is a critical insight into the metabolic regulation of tRNA modification, which opens a new field in life science. I anticipate seeing other instances of RNA modifications dynamically regulated by sensing cellular metabolic status in the coming years.”
Papers
Huan Lin, Kenjyo Miyauchi, Tai Harada, Ryo Okita, Eri Takeshita, Hirofumi Komaki, Kaoru Fujioka, Hideki Yagasaki, Yu-ichi Goto, Kaori Yanaka, Shinichi Nakagawa, Yuriko Sakaguchi, Tsutomu Suzuki, "CO2-sensitive tRNA modification associated with human mitochondrial disease," Nature Communications volume 9, Article number: 1875 (2018): May 14, 2018, doi:10.1038/s41467-018-04250-4.
Link (Publication )
)
Related links
- Graduate School of Engineering

- Department of Chemistry and Biotechnology, Graduate School of Engineering

- Suzuki Laboratory, Department of Chemistry and Biotechnology, Graduate School of Engineering






