How cells translate genetic information Chemical modifications give tRNA its ability to wobble
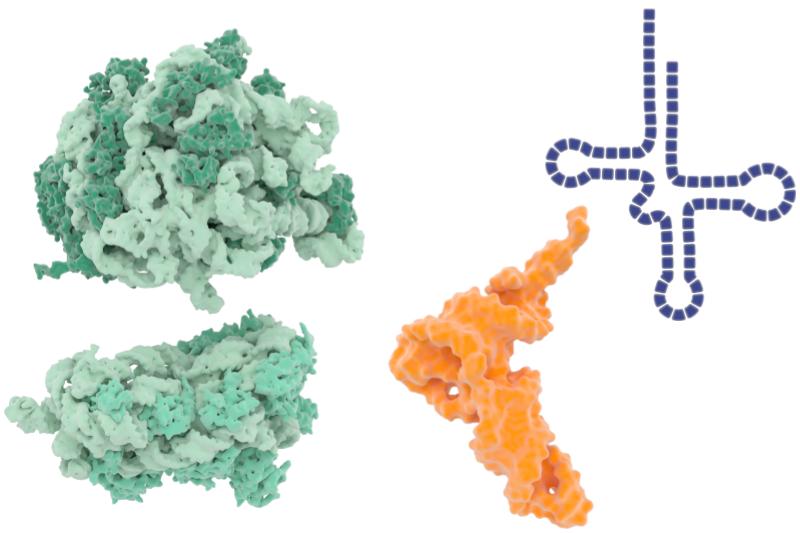
Researchers have identified another part of the machinery that cells use to efficiently and precisely turn DNA into protein. The two factors that were identified might be useful drug targets for new ways to treat bacterial infections.
The genetic information coded in our DNA goes through multiple processing steps before it becomes a functional protein. One of the final steps is translating a molecule called messenger RNA (mRNA) into a chain of amino acids, the building blocks of proteins. The translation relies on an adaptor molecule called transfer RNA (tRNA).
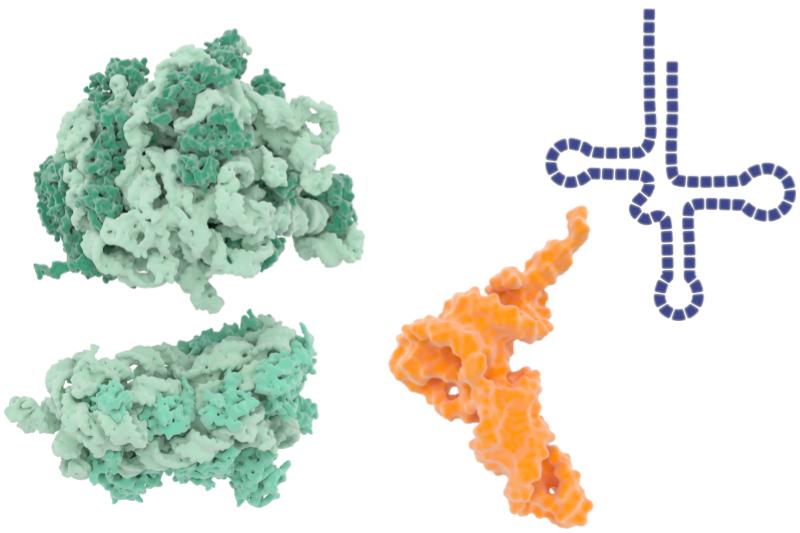
The two subunits of the ribosome are shown in green. A generic molecule of tRNA is shown in both space-filling (orange) and line drawing (blue) illustrations. Library of Science and Medical Illustrations, CC-BY
The tRNA reads the long chain of mRNA code three letters at a time to decipher which amino acid should come next in the chain. The third letter does not need to be an exact match between the mRNA and tRNA. This flexibility is known as wobble base pairing and likely keeps the translation step efficient and accurate.
Researchers at the University of Tokyo studied the properties of tRNA that make wobble base pairing possible.
"The tRNAs are decorated with various chemical modifications that play critical roles in deciphering the genetic code. We aimed to reveal how nature attains its efficient and precise system to decipher genetic information," said Professor Tsutomu Suzuki of the Department of Chemistry and Biotechnology, and leader of the laboratory where the research was performed.
RNA molecules contain more than 160 types of chemical modifications that are introduced after the RNA is transcribed from DNA.
5-carboxymethoxyuridine (cmo5U) is one important modification required for tRNA function.
"In this study, we were trying to identify novel factors responsible for the initial step of cmo5U biogenesis," said Suzuki.
The initial step in cmo5U biogenesis involves adding oxygen and hydrogen, known together as a hydroxyl group, to the tRNA molecule.
The research team identified two factors for tRNA hydroxylation that are redundantly responsible for the initial step of cmo5U biogenesis. Two enzymes, TrhP and TrhO, are involved in different chemical pathways leading to tRNA hydroxylation — prephenate-dependent and oxygen-dependent pathways, respectively.
To understand how tRNA hydroxylation happens, the research team needed to recreate tRNA modification artificially under laboratory conditions, but this had been a challenging task.
"I will never forget the day when I got a clear result showing that TrhO interacts with tRNA directly. This was a breakthrough and I succeeded in reconstituting tRNA hydroxylation the following week," said Yusuke Sakai, a doctoral student in the Suzuki lab at the time of the research and co-first author of the publication.
Researchers compared the genome of E. coli bacteria to other organisms to try to identify the gene responsible for the TrhO enzyme.
"I narrowed down the candidates to about 100 E. coli genes, and finally found a particularly suspicious gene as a target. I clearly remember the excitement when I identified the trhO gene," said Satoshi Kimura, a postdoctoral researcher in the Suzuki lab at the time of the research and co-first author of the publication.
Bacteria that were genetically modified not to have the normal cmo5U biogenesis abilities grew more slowly at certain temperatures. Other tests of modified bacteria revealed that they were less efficient at reading certain combinations of the three-letter units of mRNA, indicating their wobble base pairing abilities were impeded.
This more detailed understanding of the chemical modifications that give tRNA its wobble base pairing abilities and other functions will allow researchers to better understand the fundamental process of how DNA becomes protein.
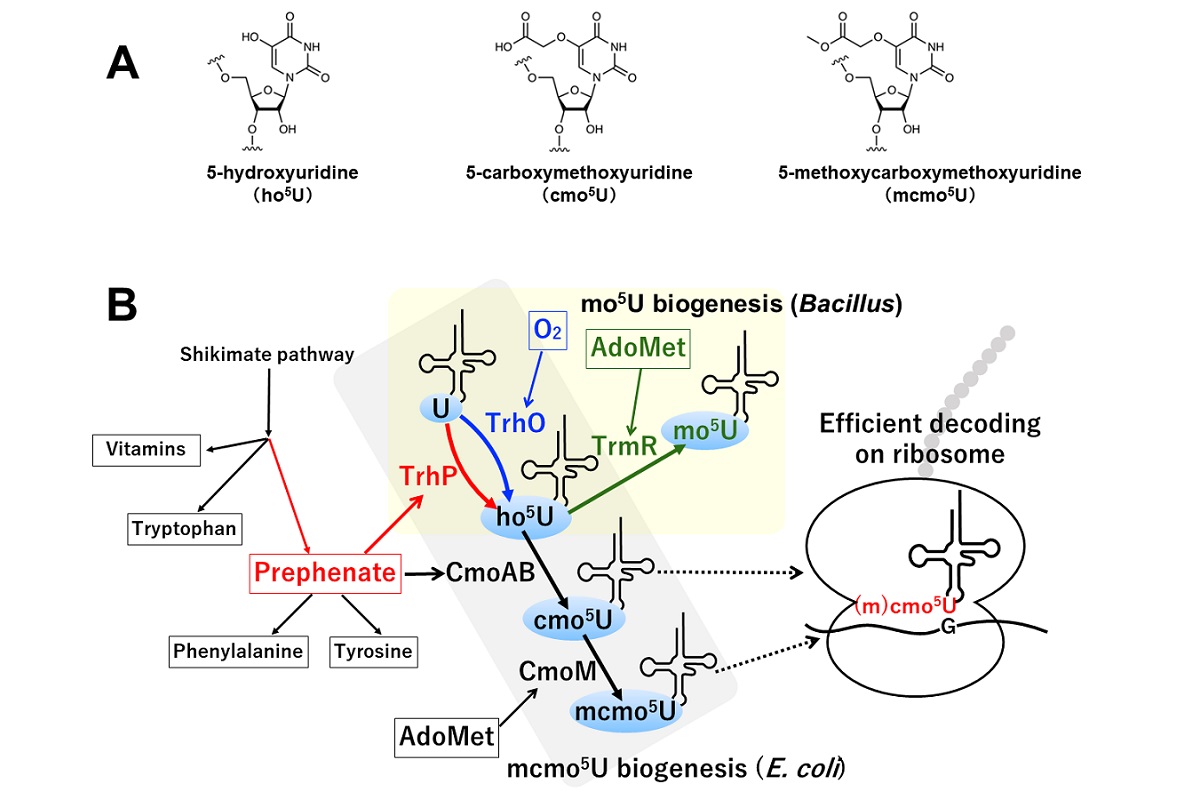
Scheme of cmo5U biogenesis initiated by tRNA hydroxylation. (A) Chemical structure of modified RNAs investigated in the research. cmo5U is derived from ho5U, while mcmo5U is a methylated derivative of cmo5U. (B) ho5U is synthesized by dual pathways. TrhP-mediated pathway requires a metabolite of shikimate pathway, while TrhO utilizes oxygen in the air. In Bacillus subtilis, ho5U is methylated by TrmR to generate mo5U, while CmoA and CmoB cooperatively introduce a carboxymethyl group to ho5U to synthesize cmo5U in E. coli, followed by further methylation to form mcmo5U. Either cmo5U or mcmo5U in tRNA plays a critical role for deciphering genetic code on the ribosome. Image by Yusuke Sakai, Satoshi Kimura, Tsutomu Suzuki
Papers
Yusuke Sakai, Satoshi Kimura, Tsutomu Suzuki, "Dual pathways of tRNA hydroxylation ensure efficient translation by expanding decoding capability," Nature Communications: July 26, 2019, doi:10.1038/s41467-019-10750-8.
Link (Publication )
)





