Small molecule's unsuspected involvement in RNA modification Discovery of tRNA-modifying enzyme that utilizes an acetate ion
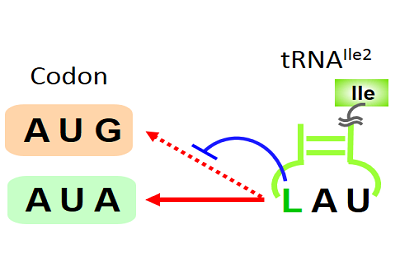
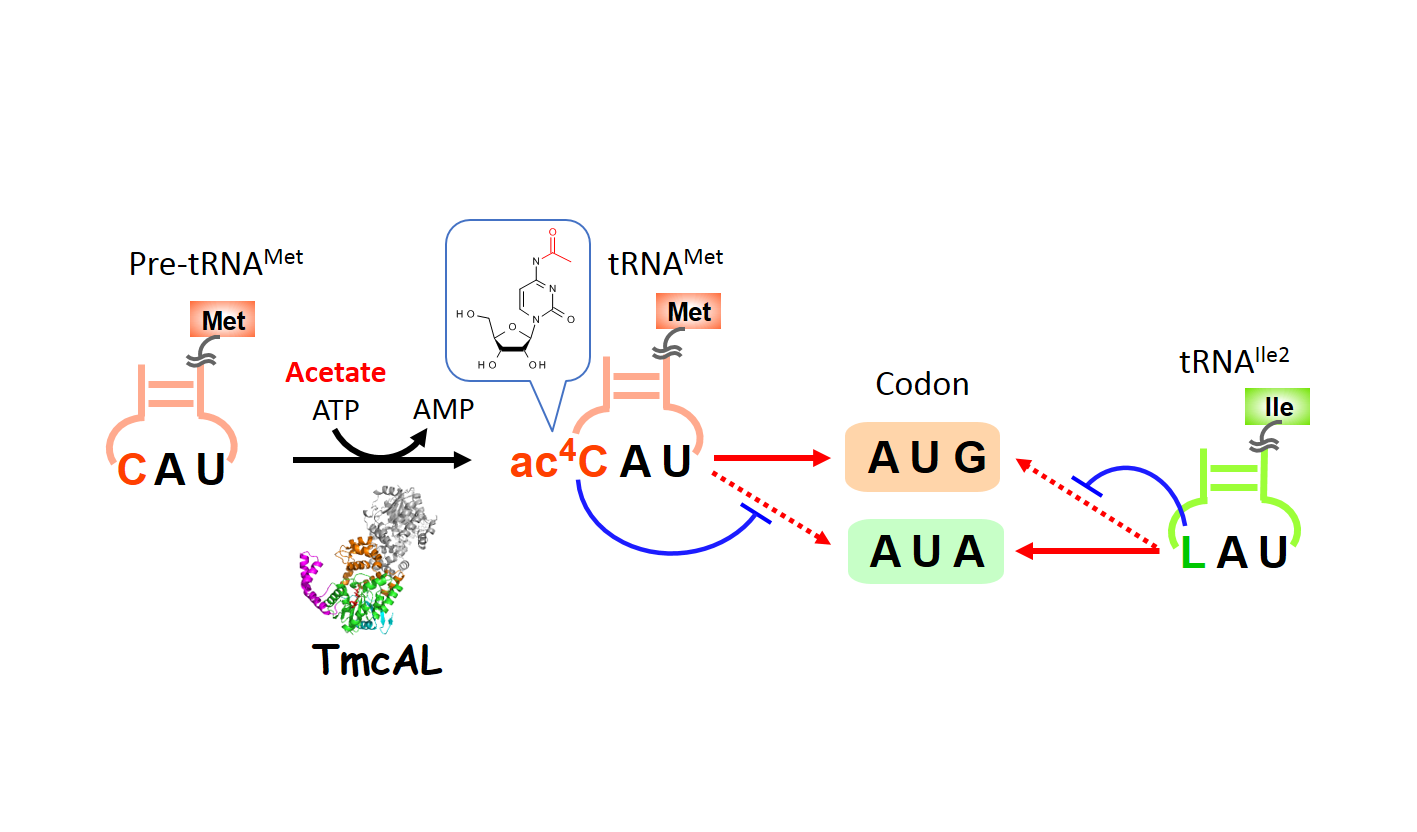
In a group of bacteria including Bacillus subtilis, TmcAL catalyzes ac4C formation in the anticodon of tRNAMet utilizing an acetate ion as a substrate. The ac4C modification cooperates with lysidine (L) modification in tRNAIle2, and plays a critical role in discriminating AUG codon (for Met) from AUA codon (for Ile), so as to maintain translational fidelity.
© 2018 Tsutomu Suzuki.
While researching bacterial RNA modification, a University of Tokyo research group identified a novel enzyme that introduces an acetyl group and chemically modifies transfer RNA, using an acetate ion as a substrate.
All living organisms synthesize proteins based on universal genetic codes. tRNA is an essential biomolecule for deciphering the genetic codes on nucleic acids. In Escherichia coli, a bacteria species found in animal intestines, tRNAMet – the tRNA responsible for methionine, an amino acid – contains N4-acetylcytidine (ac4C) as a post-transcriptional modification in its anticodon that decodes genetic information.
According to a previous study by the group, the ac4C modification is introduced by a tRNA-modifying enzyme called TmcA that uses acetyl coenzyme A (acetyl-CoA) as an acetyl donor. However, in the other subset of bacteria including Bacillus subtilis, an organism used in research that is found in soil and the gastrointestinal tracts of animals, no ac4C modification and no TmcA homolog, or similar enzyme, was reported.
The research group led by then-graduate student Takaaki Taniguchi, Project Researcher Kenjyo Miyauchi and Professor Tsutomu Suzuki of the Department of Chemistry and Biotechnology at the Graduate School of Engineering first revealed that ac4C is also present in B. subtilis tRNAMet. Then, they searched for the enzyme responsible for this modification, and identified a novel enzyme, TmcAL, responsible for the ac4C modification of B. subtilis. TmcAL belongs to an enzyme family distinct from TmcA. Surprisingly, unlike other enzymes that transfer acetyl groups, known as acetyltransferases, including TmcA, TmcAL does not use acetyl-CoA, but an acetate ion as an acetyl donor to introduce ac4C modification on tRNA. This finding indicates that ac4C modification is acquired by distinct mechanisms with different classes of enzymes, implying it is established by convergent evolution among bacterial phyla, as large subdivisions are called. Moreover, the research group solved the crystal structure of TmcAL and revealed the molecular basis of how TmcAL activates an acetate ion with adenylation, a process that modifies proteins, and catalyzes ac4C formation.
The research group also found a strong genetic interaction between the tmcAL gene and the tilS gene that encodes the enzyme responsible for synthesizing lysidine in tRNAIle to facilitate decoding of isoleucine codon, suggesting that ac4C and lysidine act cooperatively to maintain accurate protein synthesis. In fact, they found evidence clearly showing that the genetic code for isoleucine, an amino acid, was drastically misdecoded by methionine in the absence of tmcAL under tilS repression, revealing that ac4C modification plays a critical role in accurate decoding of genetic information.
The findings obtained by the research group provide insights that connect RNA research with metabolic study. This research also contributes to our understanding of how organisms synthesize proteins precisely by elucidating the accurate decoding mechanism mediated by tRNA modifications.
"This research started when we accidentally found ac4C modification in an organism not bearing TmcA," says Taniguchi. He continues, "I'm excited to discover TmcAL, and its catalytic mechanism for ac4C formation using an acetate ion as a substrate. Many questions related to RNA modifications still remain to be investigated. I hope my research contributes to future studies on RNA modifications."
Papers
Takaaki Taniguchi, Kenjyo Miyauchi, Yuriko Sakaguchi, Seisuke Yamashita, Akiko Soma, Kozo Tomita, Tsutomu Suzuki, "Acetate-dependent tRNA acetylation required for decoding fidelity in protein synthesis," Nature Chemical Biology: August 27, 2018, doi:10.1038/s41589-018-0119-z.
Link (Publication )
)
Related links
- Graduate School of Engineering

- Department of Chemistry and Biotechnology, Graduate School of Engineering

- Suzuki Laboratory, Department of Chemistry and Biotechnology, Graduate School of Engineering






