A new concept for restriction enzymes A DNA glycosylase can act as a restriction enzyme

The genetic modification of DNA sequences is used in the mass production of medicines and genetic therapy to introduce corrected sequences into patient genomes. This technology depends on the ability of restriction enzymes to recognize and cleave DNA strands at specific nucleotide sequences. It was thought that restriction enzymes are sequence-specific endonucleases that cut DNA strands by cleaving the connection between the sugar and phosphate portions (phosphodiester bonds) in the backbone of DNA.
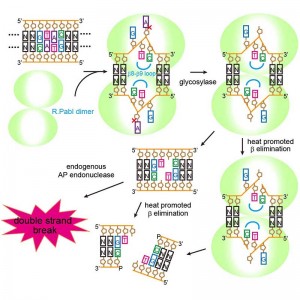
© Masaru Tanokura. R.PabI recognizes a 5’-GTAC-3’ sequence and produces two opposing apurinic/apyrimidinic (AP) sites. The opposing AP sites are cleaved by heat-promoted β elimination, and/or by endogenous AP endonucleases of host cells to introduce a double-strand break.
By combining X-ray crystal structure analysis with a variety of analytical techniques, a research group led by University of Tokyo Graduate School of Agricultural and Life Sciences Professor Masaru Tanokura has found that the restriction enzyme R.PabI from the hyperthermophilic archaea Pyrococcus abyssi acts as a DNA glycosylase, cutting the bond connecting the base to the deoxyribose sugar of the DNA backbone. R.PabI catalyzes the cleavage between the adenine base and the sugar portion of DNA in an identified sequence, generating what is termed an abasic site. This abasic site is then cleaved by heat-promoted β elimination and/or the action of other enzymes, disconnecting the double-stranded DNA.
This discovery of a mechanism in bacterial cells to recognize a specific sequence and cleave DNA at that site overturns accepted knowledge. This is the first discovery of a restriction enzyme with DNA glycosylase activity, but it is expected that more will be found.
This research was published in the January 24 issue of the British science journal Nature Communications.
Press release (Japanese)
Paper
Ken-ichi Miyazono, Yoshikazu Furuta, Miki Watanabe-Matsui, Takuya Miyakawa, Tomoko Ito, Ichizo Kobayashi and Masaru Tanokura,
“A sequence-specific DNA glycosylase mediates restriction-modification in Pyrococcus abyssi”,
Nature Communications Online Edition: 2014/1/24, doi: 10.1038/ncomms4178.
Article link
Links
Graduate School of Agricultural and Life Sciences
Department of Applied Biological Chemistry, Graduate School of Agricultural and Life Sciences






